Proteins Mosaic Q Project
from 01/01/2026 until 01/01/2030
The Proteins Mosaic Q Project seeks to promote research on amino acid clustering patterns in the 3D structure of proteins. Specifically, it analyzes how amino acids according cluster according to their chemical type (polar, hydrophobic, acidic, basic, special) following a specific pattern revealed by mathematical/statistical analysis of over 160,000 publicly available crystallographic structures.
The referred clustering pattern reveals that, aside from the hydrophobic core, residues of the same chemical family (polar, acidic, basic or special) tend to cluster in groups of specific size and shape in the 3D structure of proteins. This pattern is designated as Mosaic Q, since it can be quantified mathematically by a parameter named Q (see more information at https://proteins-mosaic-q.org/about/).
Importantly, the referred property (Mosaic Q) can also be assessed directly visually by observing multiple examples of images of protein structures rendered by visualization programs like Jmol. Citizens can participate by rendering their own images of proteins to check as independent observers whether the patterns appears, so that a collaborative repository is being generated.
The project has already been admitted in the international citizen science platform SciStarter, and has currently more than 330 followers on LinkedIn (mostly from the bioinformatics and data analysis fields) and other social media sites.
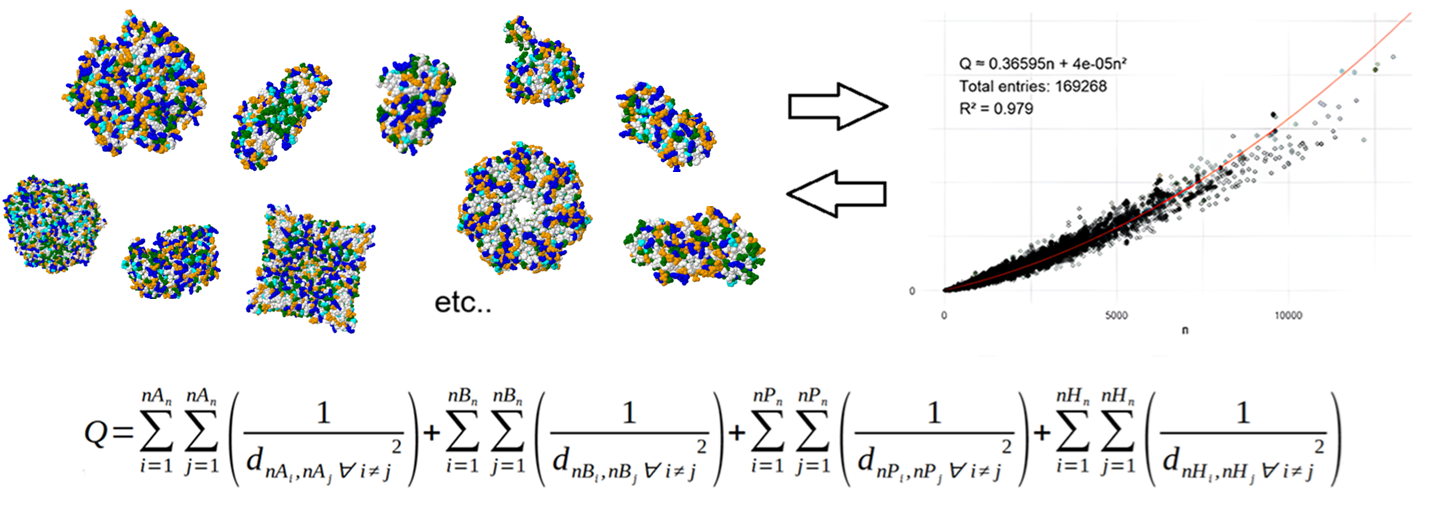
Overview of the project. Big data analysis (over 160,000 structures analyzed) reveals a characteristic clustering pattern of amino acids in the 3D structures of proteins. This is quantified by parameter Q, which is based on the calculation of euclidean distances of amino acids of the same type in the 3D structures. The mosaic pattern can also be appreciated by inspecting visually multiple examples, which is how citizens can contribute providing their own independent observations in images rendered by themselves.
Aim
The objective is to gather sufficient evidence to promote research in this field by the scientific community. Each person who participates is providing evidence to encourage research into the spatial clustering pattern of amino acids in the 3D structure of proteins according to their chemical type, since statistical analysis and multiple visualization examples point towards this pattern.
Ultimately, it aims to foster involvement of the scientific community in this field of structural biology. We believe that the future development of various protein-based drugs or drugs that have proteins as their therapeutic target could potentially benefit if this field is investigated.